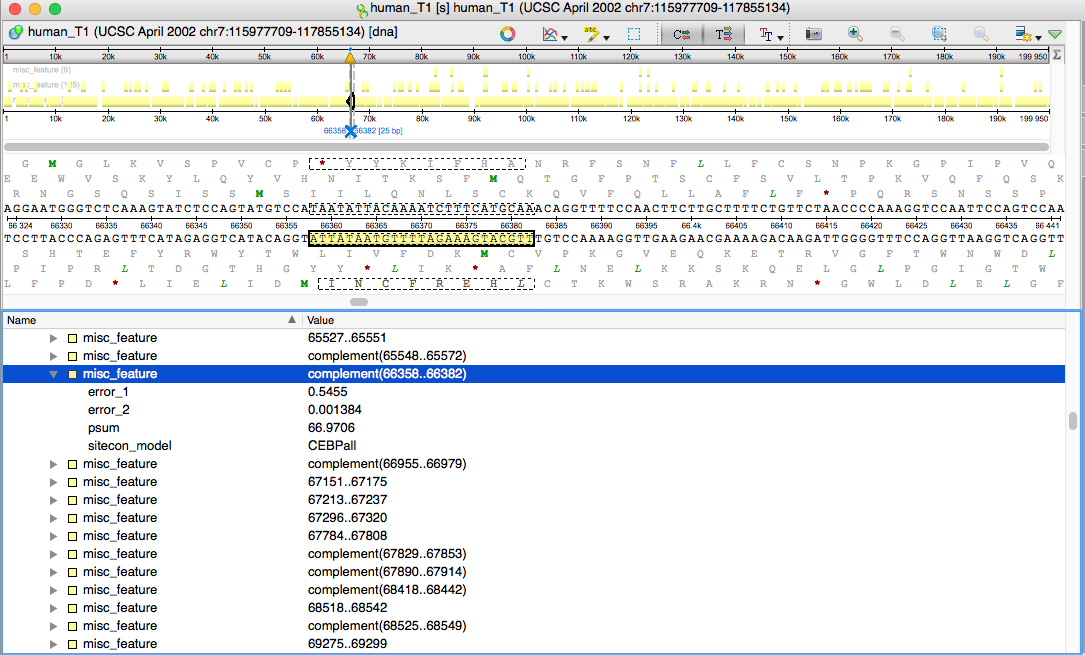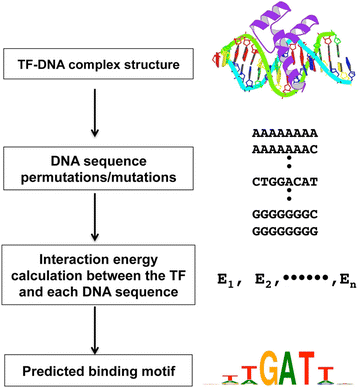
An efficient algorithm for improving structure-based prediction of transcription factor binding sites | BMC Bioinformatics | Full Text
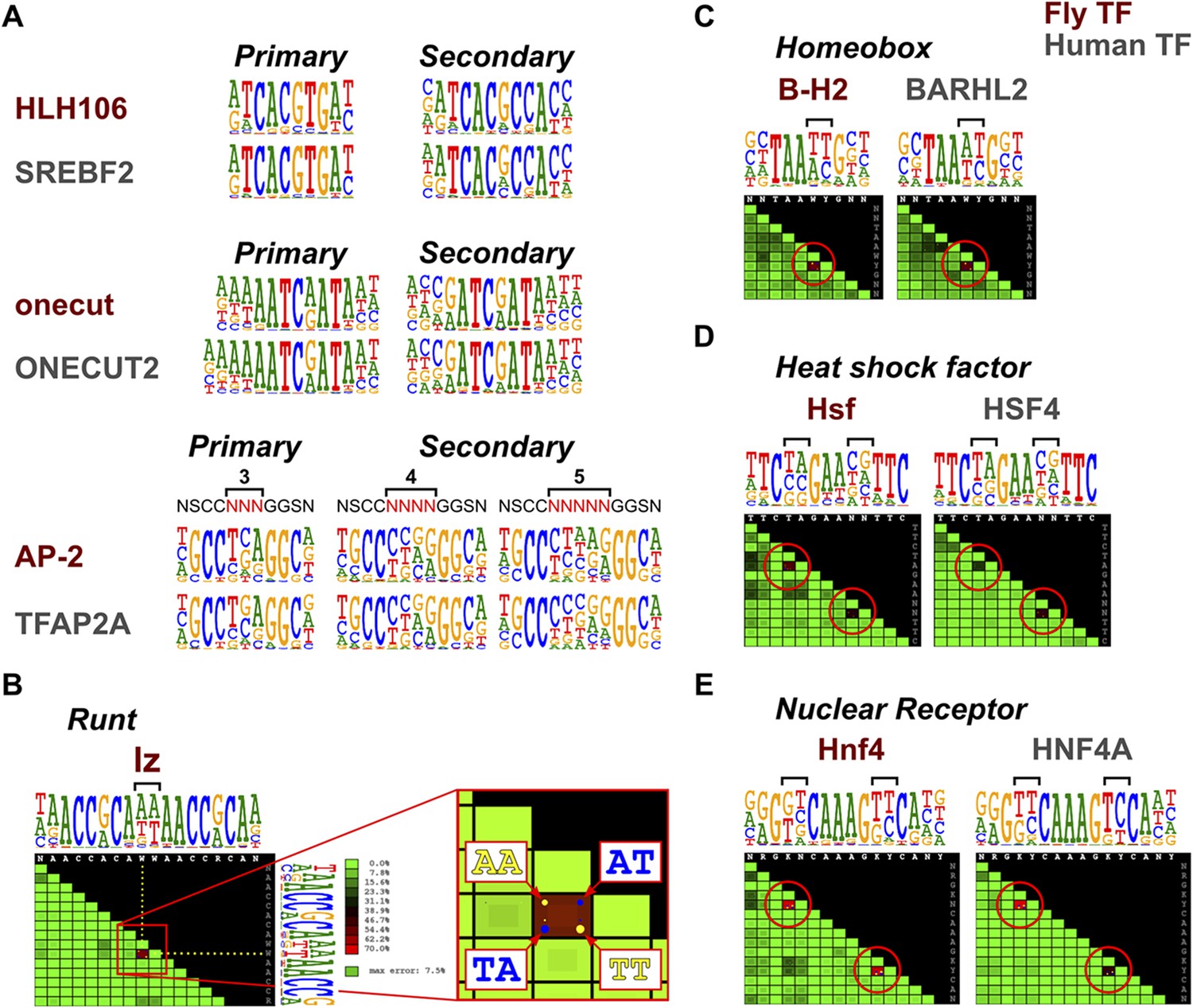
Conservation of transcription factor binding specificities across 600 million years of bilateria evolution | eLife
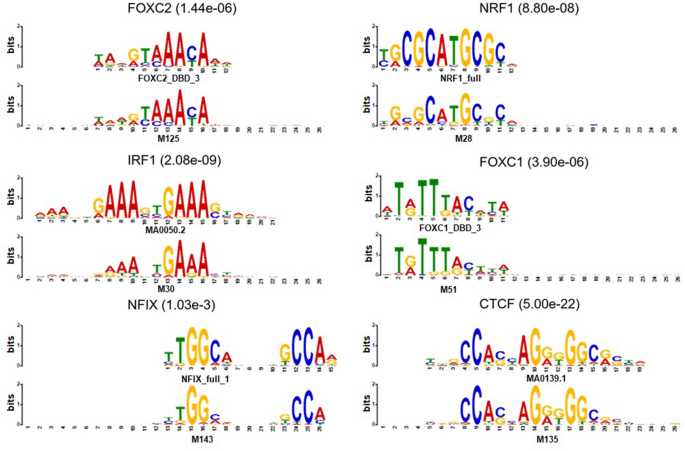
Enhancing the interpretability of transcription factor binding site prediction using attention mechanism | Scientific Reports

Leopard: fast decoding cell type-specific transcription factor binding landscape at single-nucleotide resolution | bioRxiv

Transcription factor binding sites Prediction. The sequence around the... | Download Scientific Diagram
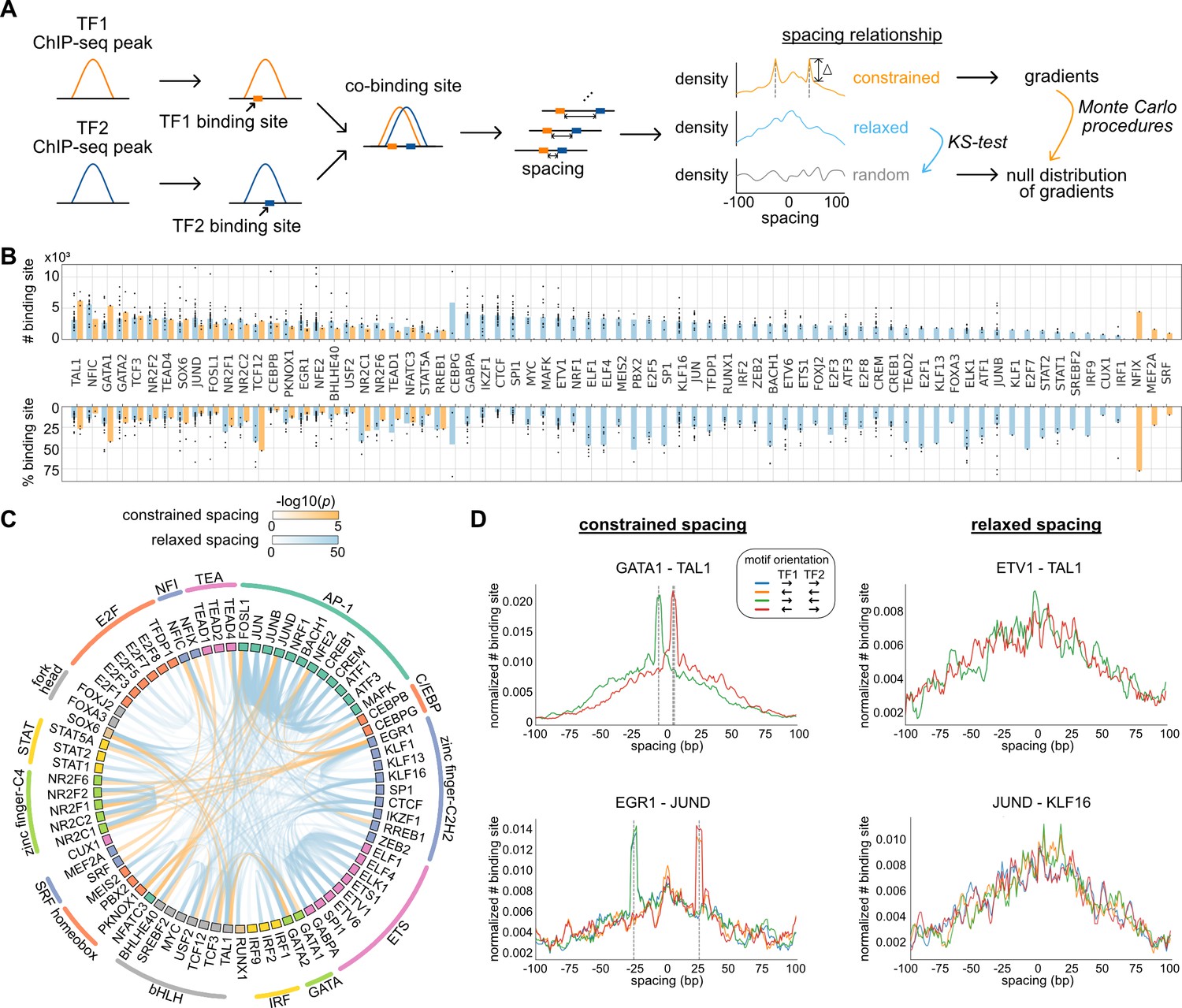
Systematic analysis of naturally occurring insertions and deletions that alter transcription factor spacing identifies tolerant and sensitive transcription factor pairs | eLife

IMAGE predicts transcription factor binding sites with high confidence.... | Download Scientific Diagram

Prediction of transcription factor bindings sites affected by SNPs located at the osteopontin promoter - ScienceDirect

Prediction of transcription factor bindings sites affected by SNPs located at the osteopontin promoter - ScienceDirect
Binding Site Graphs: A New Graph Theoretical Framework for Prediction of Transcription Factor Binding Sites | PLOS Computational Biology
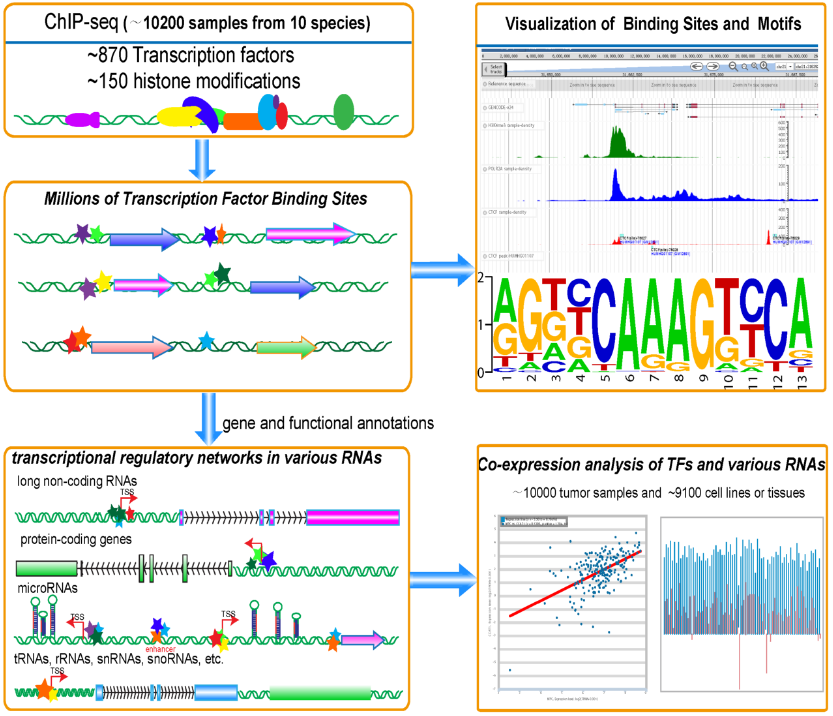
Transcription Factor Binding Sites, Motifs and Expression Profiles from ~10200 ChIP-seq and ~20000 RNA-seq samples
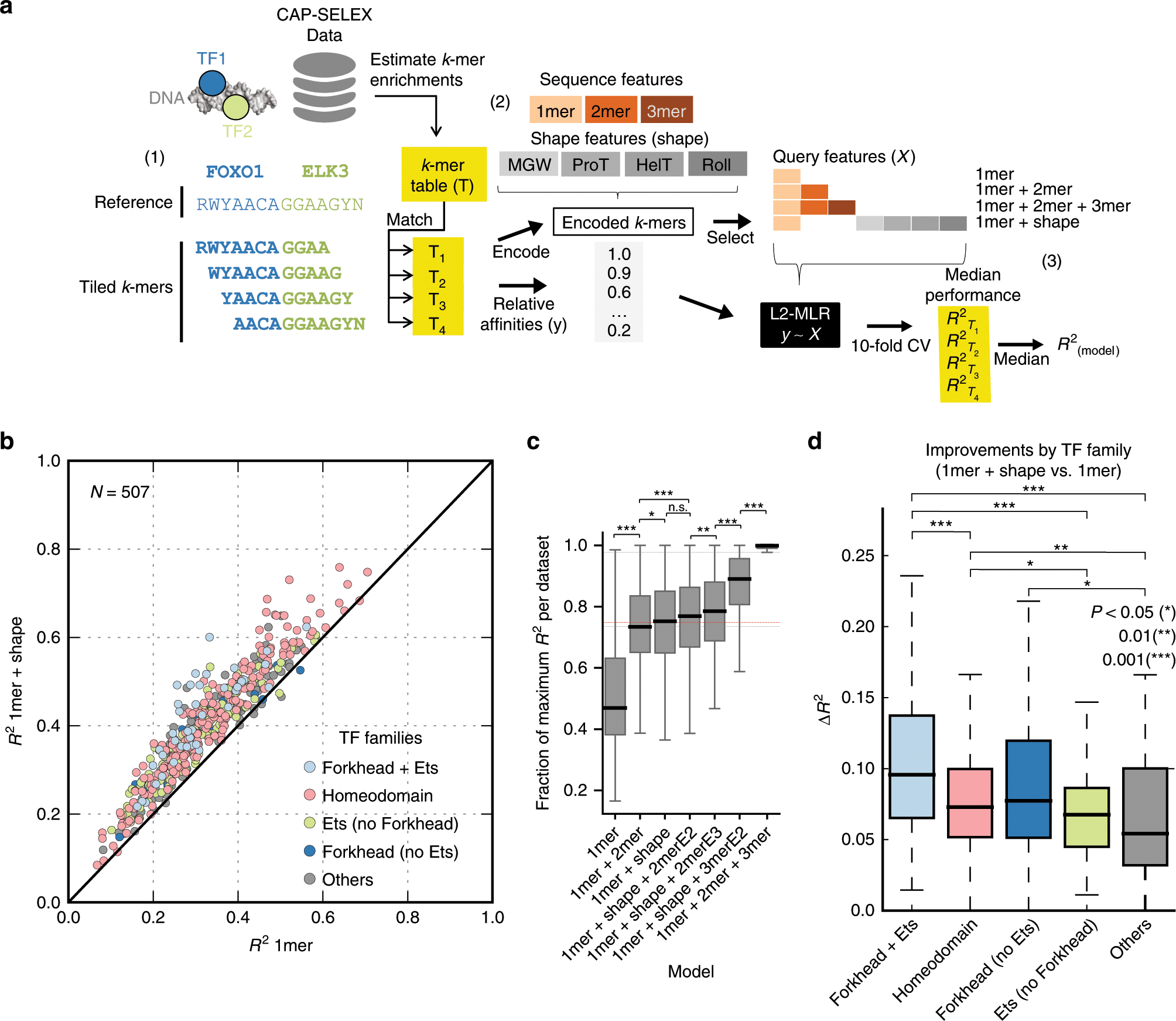
Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions | Nature Communications
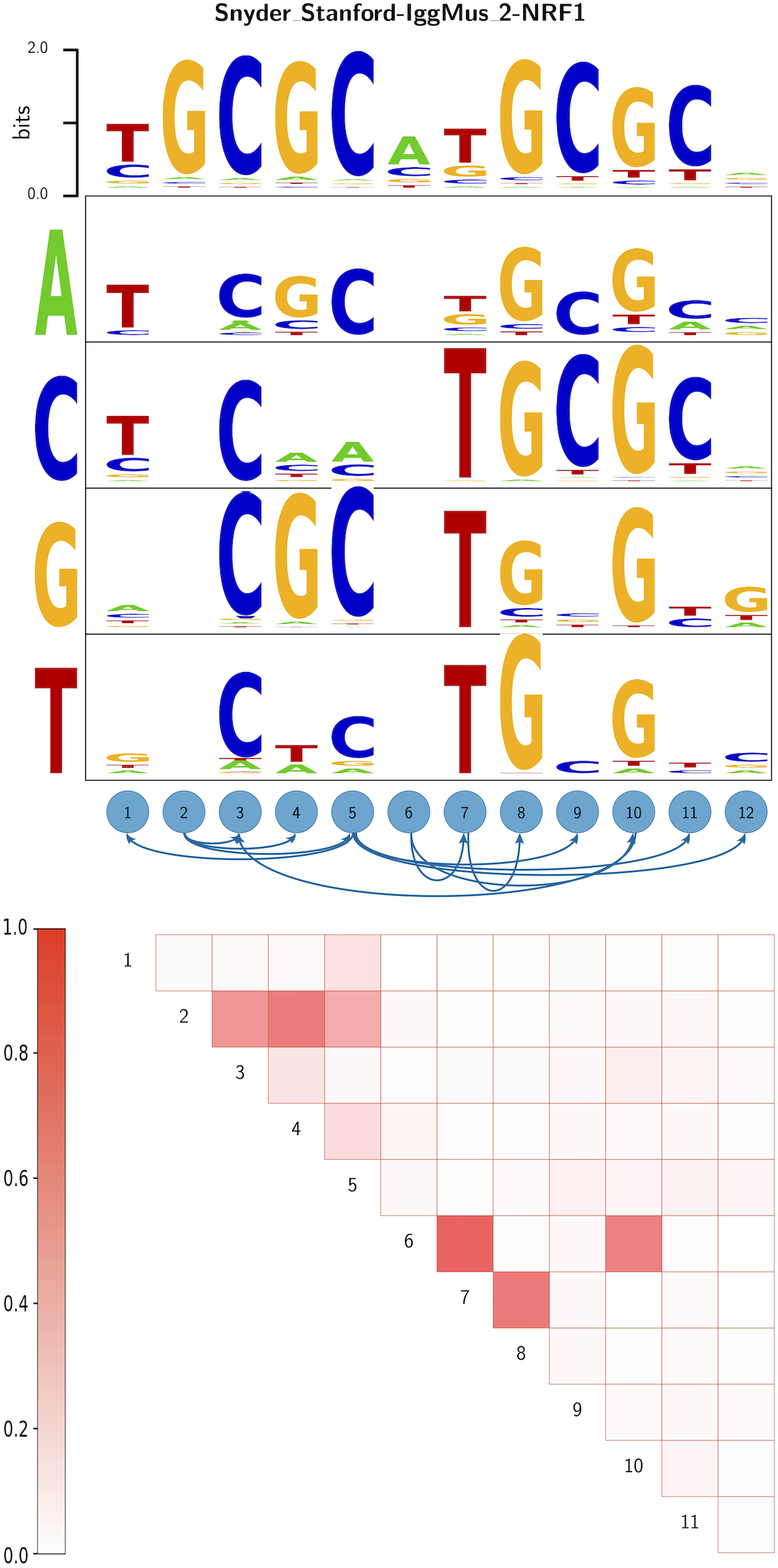
Automated incorporation of pairwise dependency in transcription factor binding site prediction using dinucleotide weight tensors | NimwegenlabNimwegenlab | Erik van Nimwegen group page