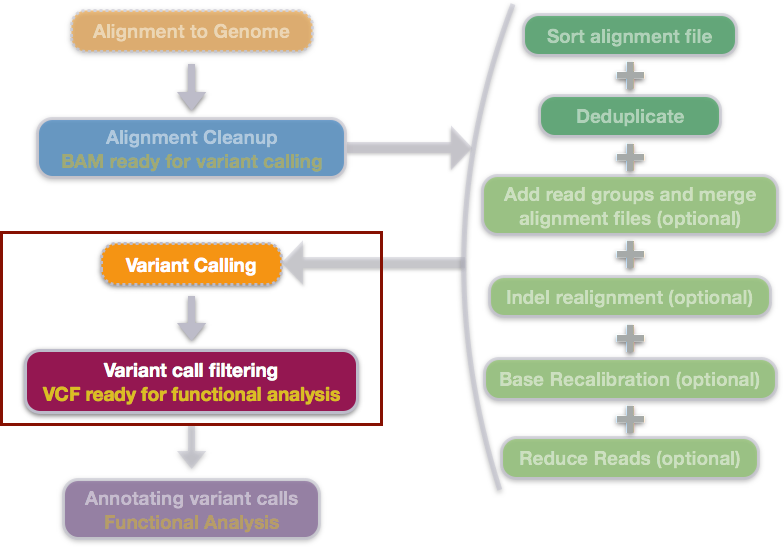
Increasing calling accuracy, coverage, and read-depth in sequence data by the use of haplotype blocks | PLOS Genetics

Long walk to genomics: History and current approaches to genome sequencing and assembly - ScienceDirect
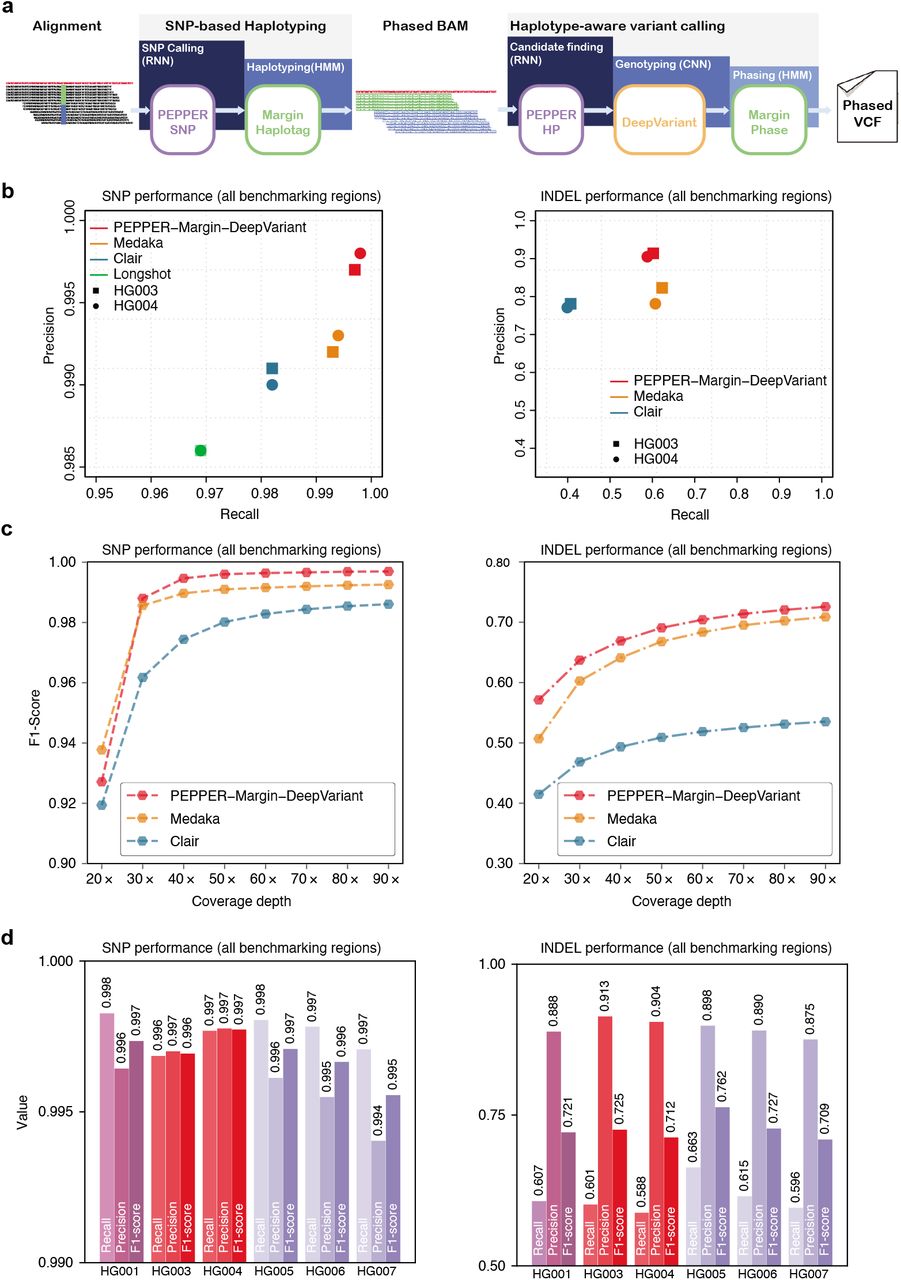
Haplotype-aware variant calling enables high accuracy in nanopore long-reads using deep neural networks | bioRxiv
Sequencing 101: ploidy, haplotypes, and phasing — how to get more from your sequencing data - PacBio
Sequencing 101: ploidy, haplotypes, and phasing — how to get more from your sequencing data - PacBio
Integrating mapping-, assembly- and haplotype-based approaches for calling variants in clinical sequencing applications | Nature Genetics

Targeted linked-read sequencing for direct haplotype phasing of maternal DMD alleles: a practical and reliable method for noninvasive prenatal diagnosis | Scientific Reports

Variant calling: Considerations, practices, and developments - Zverinova - 2022 - Human Mutation - Wiley Online Library

Genes | Free Full-Text | Inferring Signatures of Positive Selection in Whole-Genome Sequencing Data: An Overview of Haplotype-Based Methods
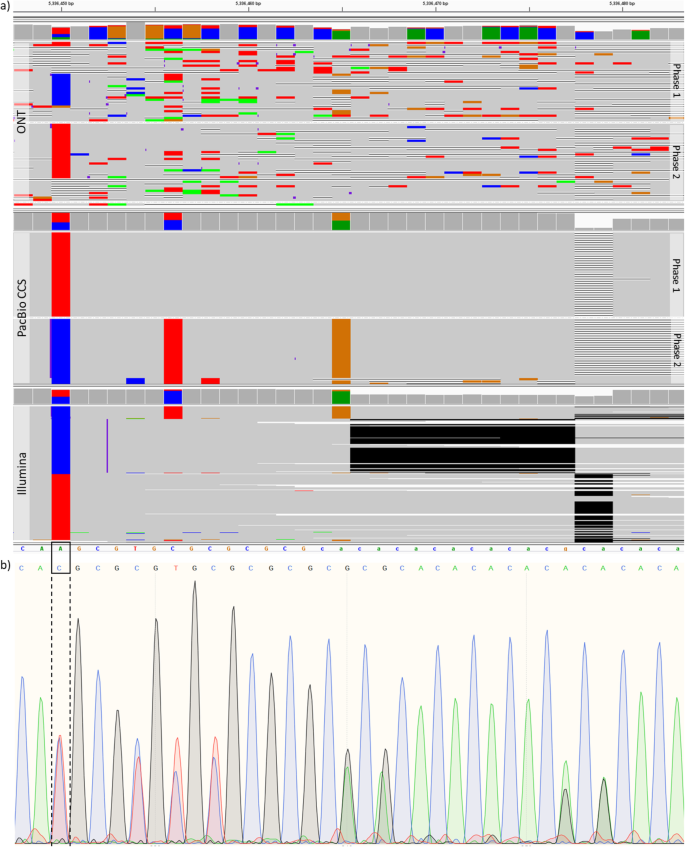
NanoCaller for accurate detection of SNPs and indels in difficult-to-map regions from long-read sequencing by haplotype-aware deep neural networks | Genome Biology | Full Text






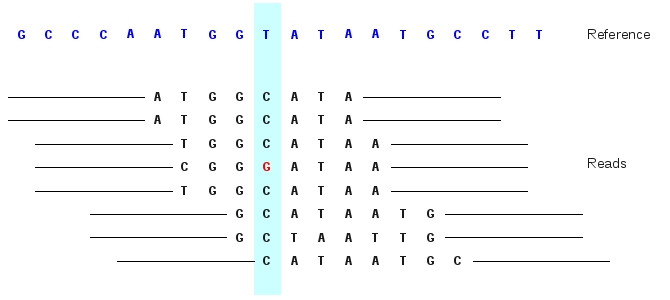
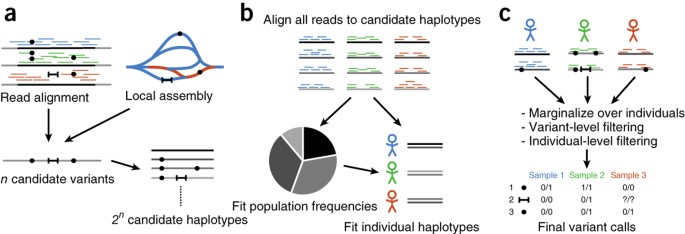
![PDF] Haplotype-based variant detection from short-read sequencing | Semantic Scholar PDF] Haplotype-based variant detection from short-read sequencing | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cc5c9c18527d3f49f0a8c16308e4aa7c67f782ad/5-Figure2-1.png)