XBSeq – Differential expression analysis of RNA sequencing data by incorporating non-exonic mapped reads | RNA-Seq Blog

Amazon.com: Differential expression analysis for sequence count data eBook : Various Authors: Kindle Store
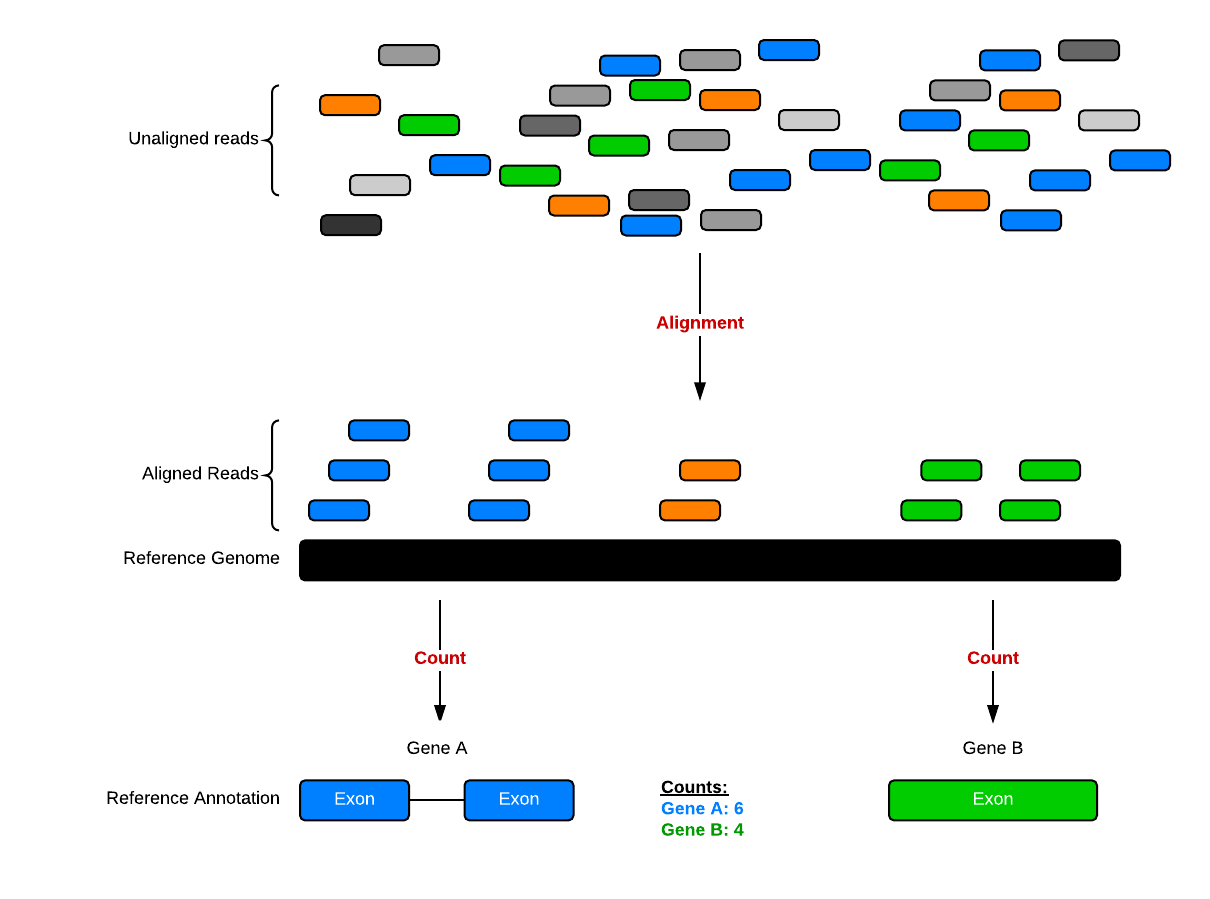
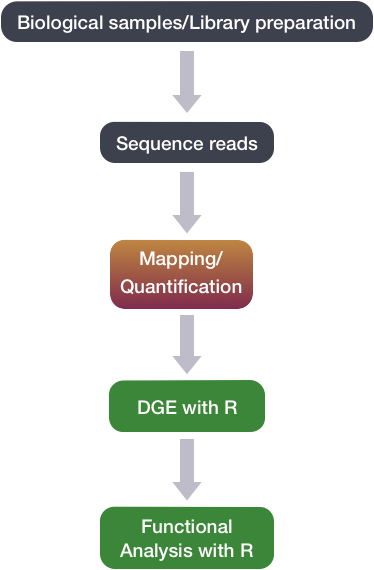
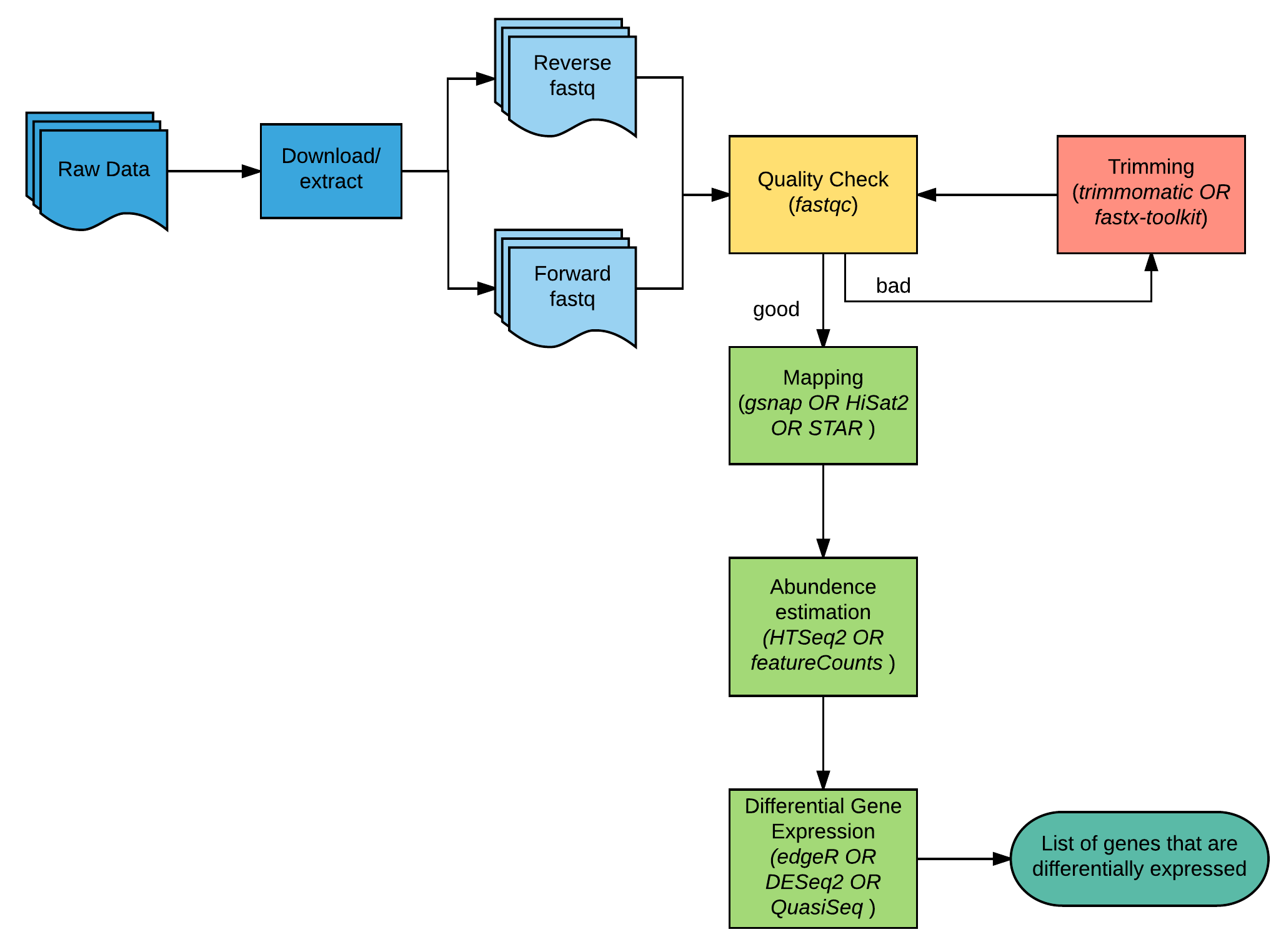
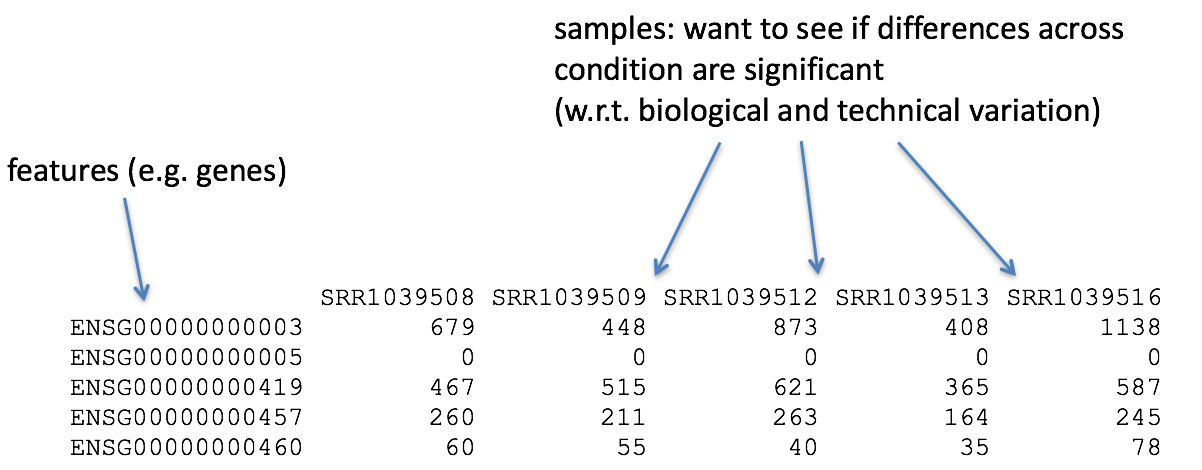
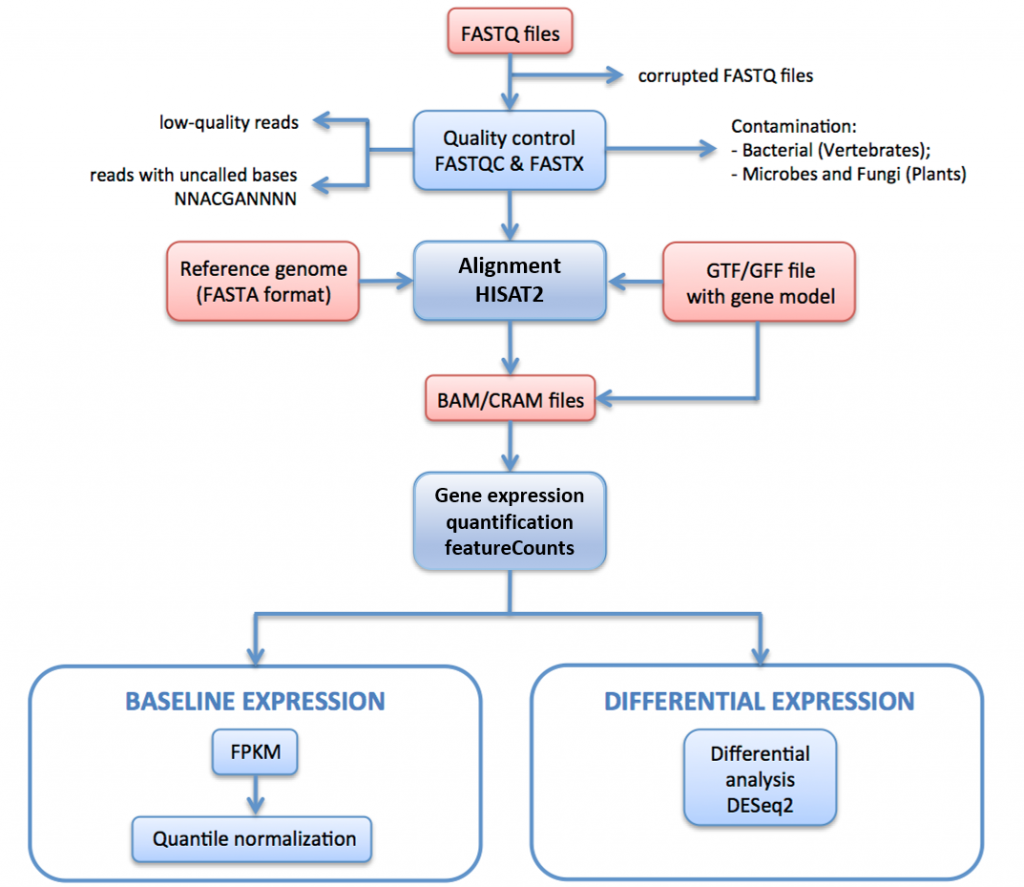
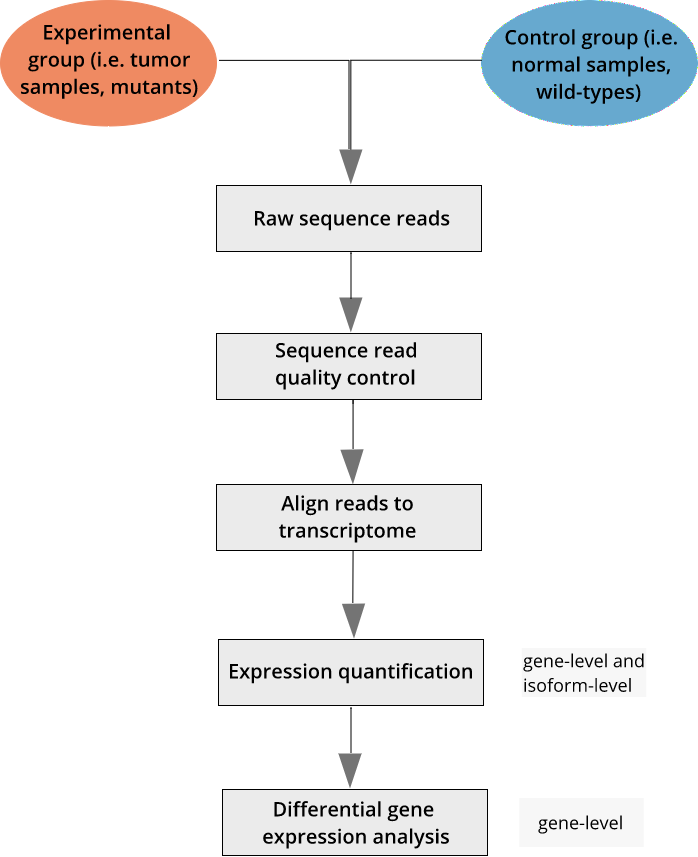
Set up and overview for gene-level differential expression analysis | Introduction to DGE - ARCHIVED

Comparison of Regression Methods to Identify Differential Expression in RNA-Sequencing Count Data from the Serial Analysis of Gene Expression
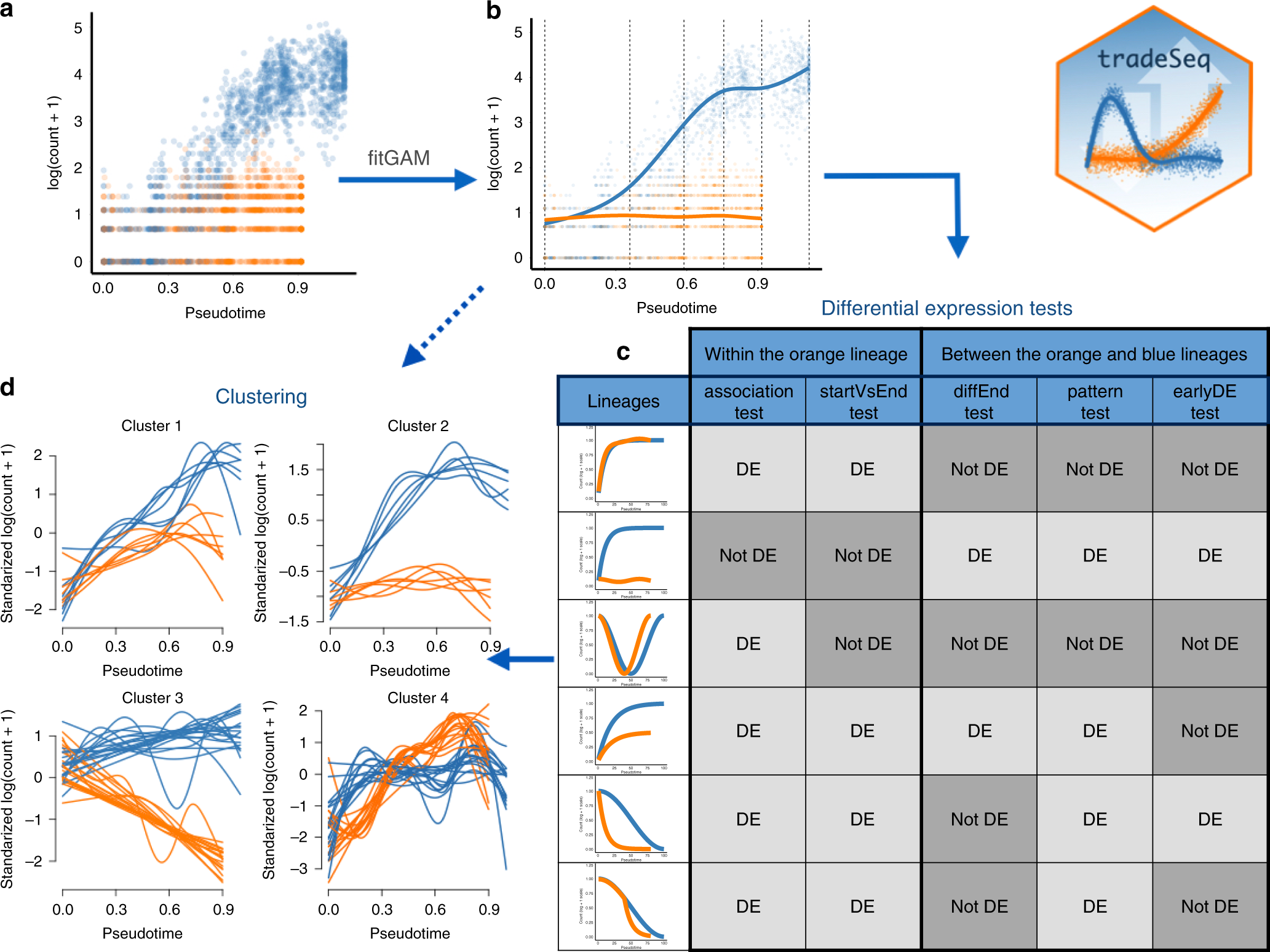
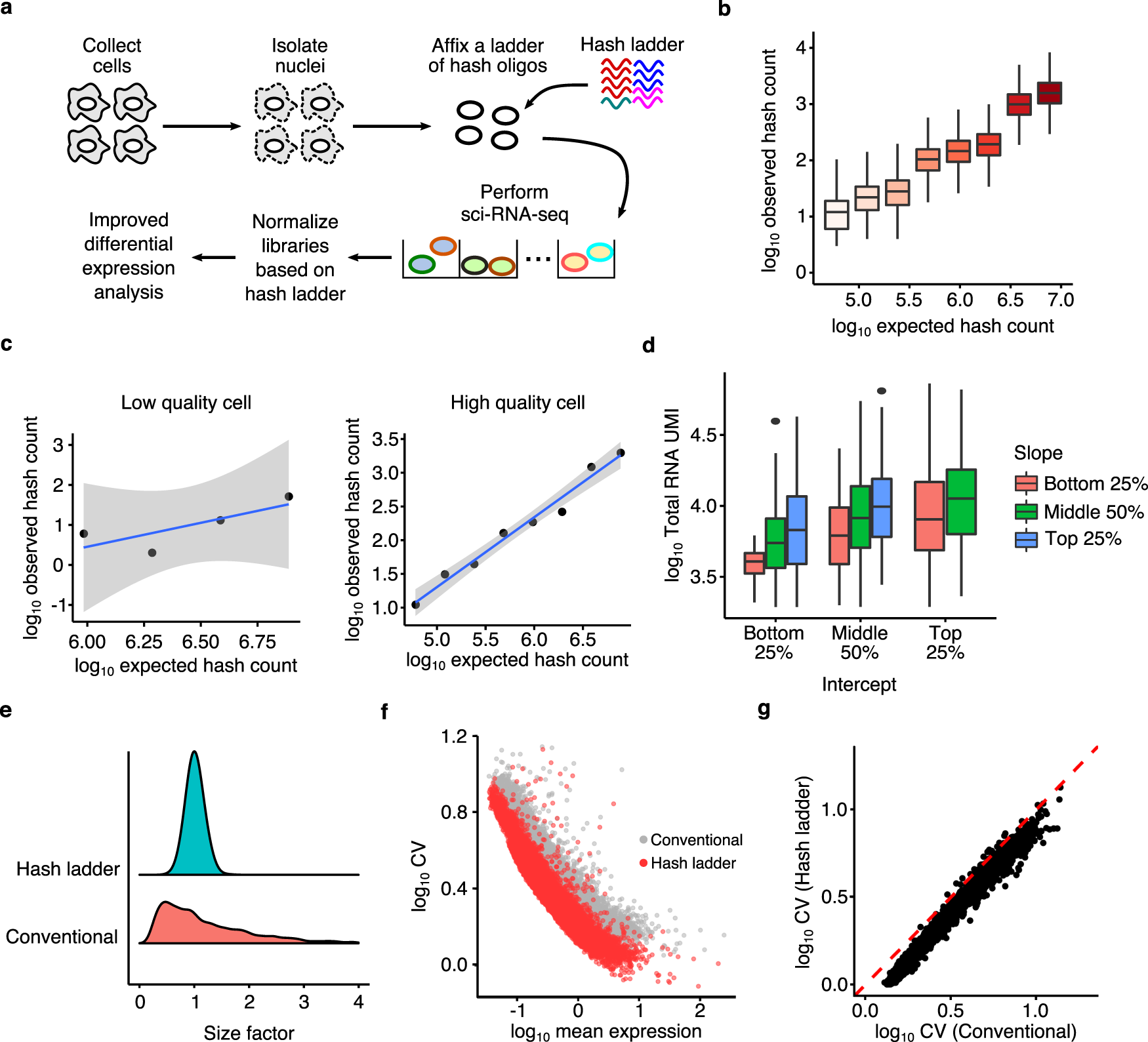
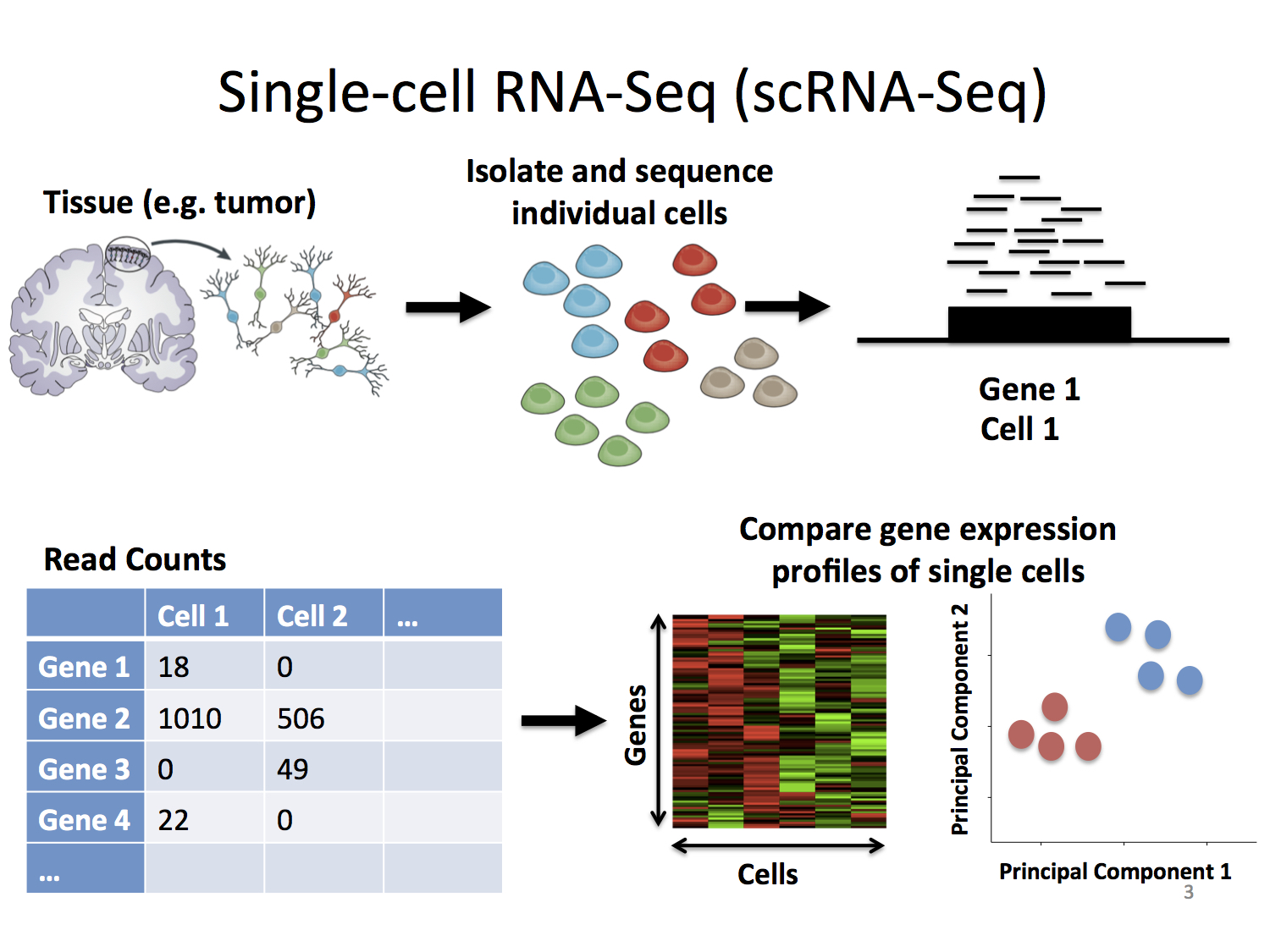
Trajectory-based differential expression analysis for single-cell sequencing data | Nature Communications
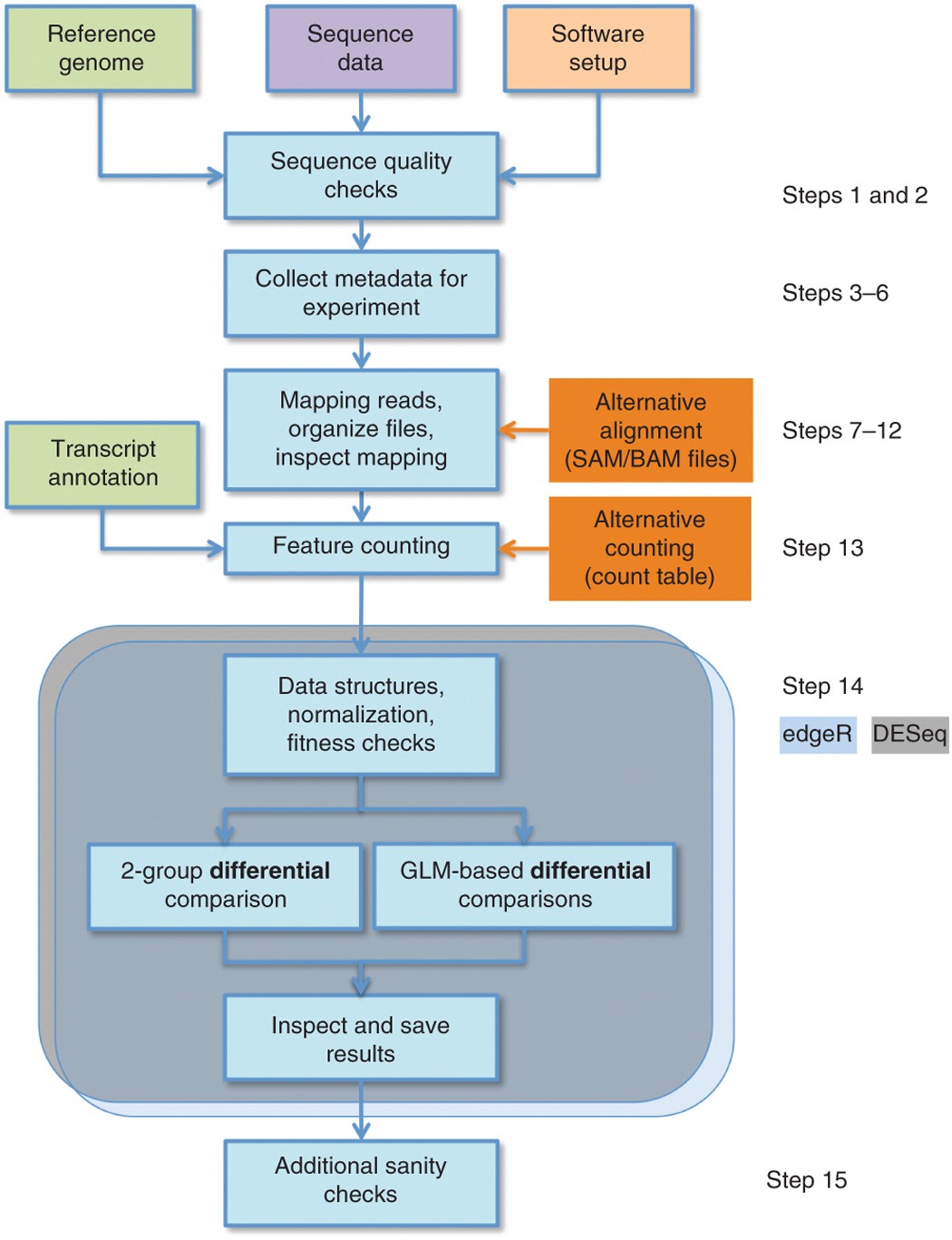
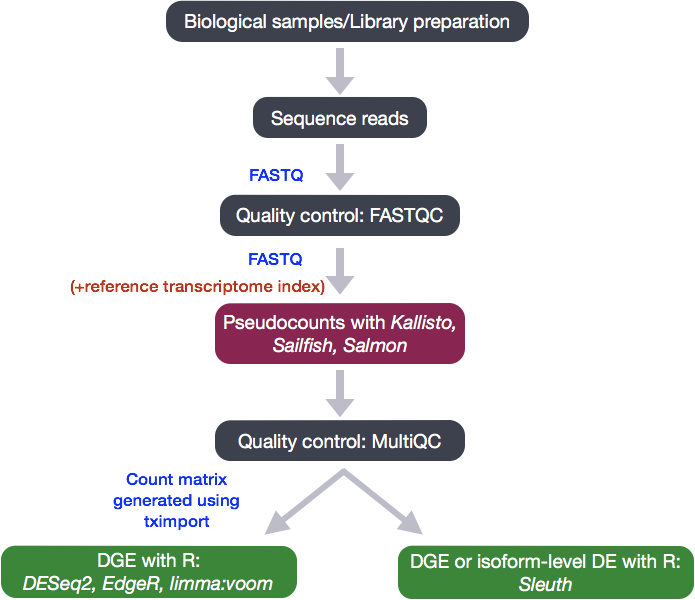
Count-based differential expression analysis of RNA sequencing data using R and Bioconductor | Nature Protocols
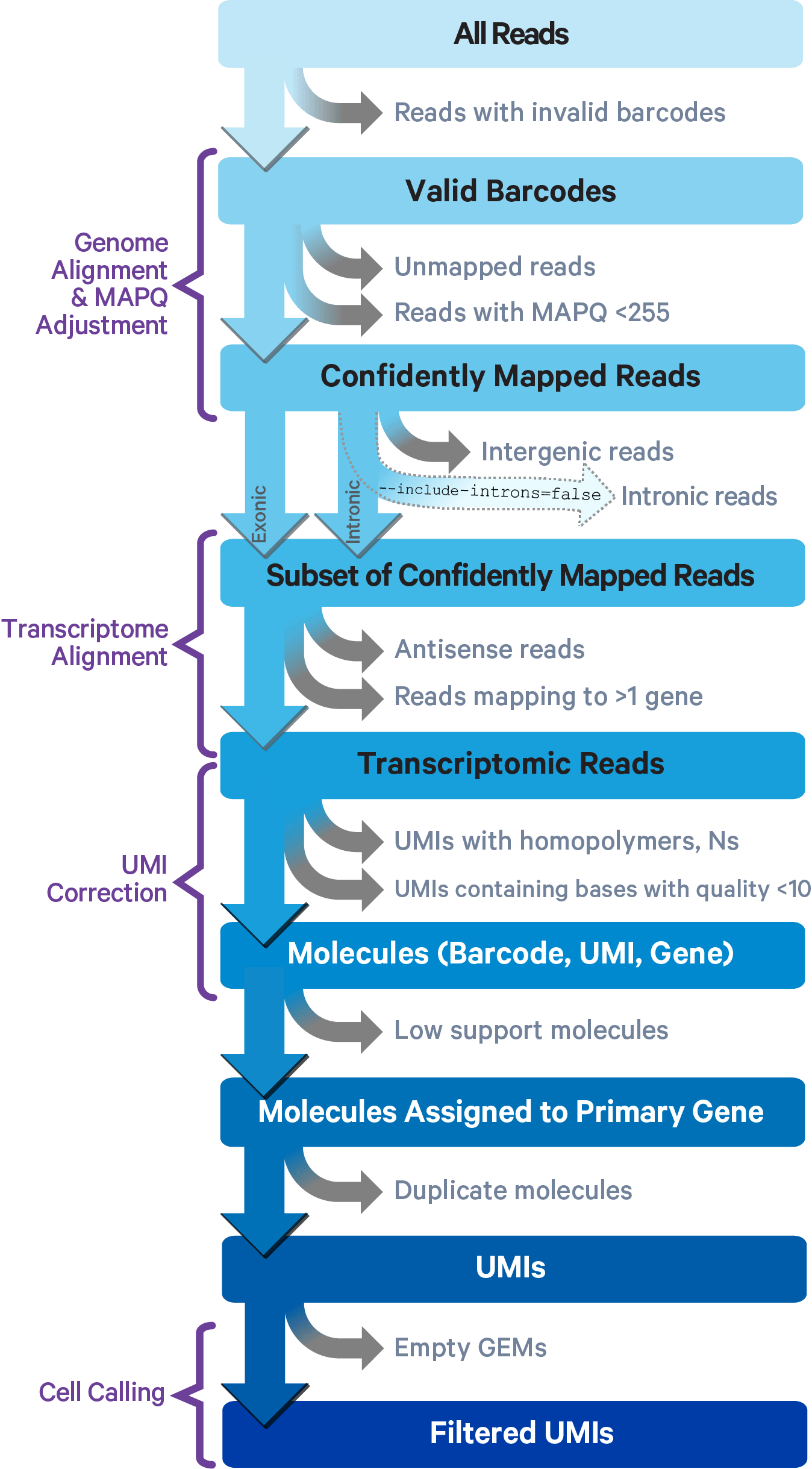
Gene Expression Algorithms Overview -Software -Single Cell Gene Expression -Official 10x Genomics Support